|
Optionally eliminate rare codons and restriction sites. New codon optimization algorithm lets you match the codon usage from your organism of choice by generating a new sequence, starting from either a protein or nucleotide sequence. The HMM alignment engine improves both quality and speed of alignments compared to the older ClustalW aligner.īetter back translate and codon optimization Scale up your alignments with Clustal Omega, now included with Geneious Prime. Longread sequence mapping with Minimap2 (external plugin)įast and easy alignment of Oxford Nanopore and PacBio data to a reference sequence using an industry recommended tool, without the hassle of the command lineĬlustal Omega replaces ClustalW for better, faster alignments Quickly visualize restriction sites affected by Dam, Dcm and EcoKI methylation. 
Highlighted methylation sites for restriction enzymes 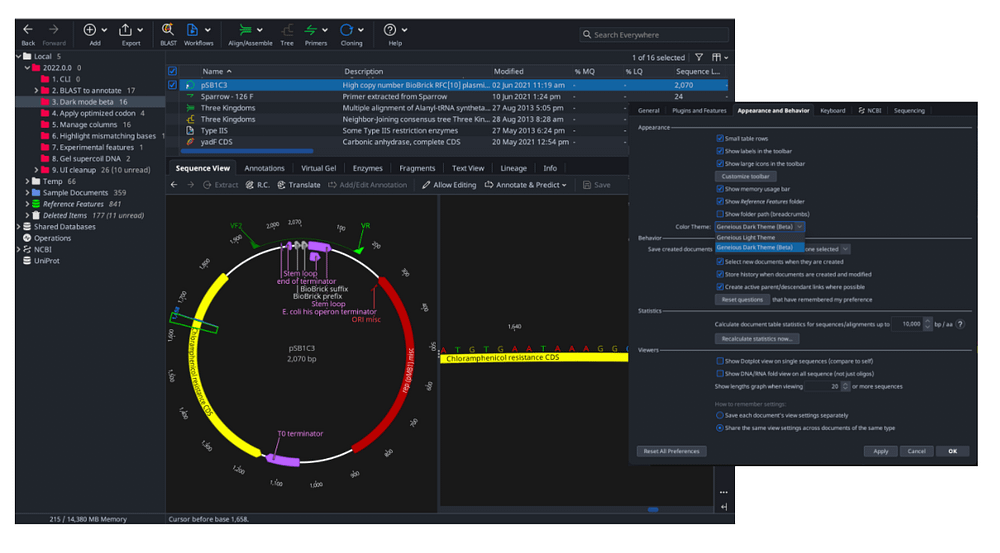
2016 CRISPR on-target scoring algorithm.Įasily sort and export your protein and nucleotide statistics within the document table. 
Enjoy the new look and feel of Geneious Prime with an updated design, including new icons and font.Ĭonfidently design your guide RNAs using the Doench et al.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
